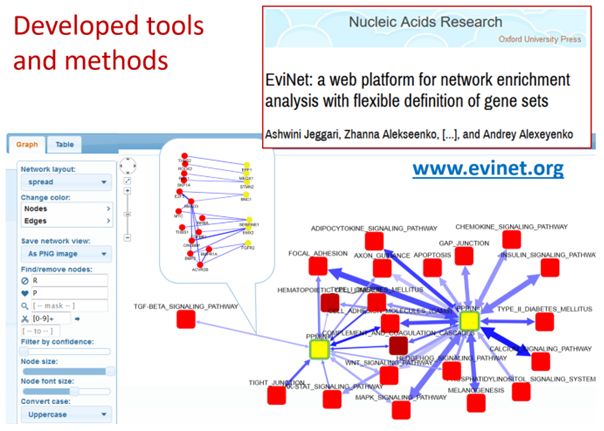
Pathway enrichment analyses are usually limited to a subset of genes (or proteins) assigned to pathways (<50%) AND covered by the data profiling (transcriptomics, variant calling, a protein panel – 1…5% of the known proteins). This limitation does not exist for our method of network enrichment analysis (NEA): indeed, practically every known gene possesses a number of network links sufficient for its functional characterization against pathways, GO terms, or even single network nodes of key importance.
The web interface of NEA can be found at https://www.evinet.org/
Coming soon:
· Robust diagnostic panels inferred via network analysis.
· The universe of (very large) public data sets: how much your research can benefit from it?
· Strategies of data integration for omics data.